Javascript array splice efficiency11/25/2023  
In that work, we used CBEs to disrupt genes by mutating CAG, CGA, and TGG codons to introduce premature stop codons (pmSTOPs, previously termed iSTOP or CRISPR-STOP) 20, 21, or by mutating the conserved splice-site motifs to disrupt RNA splicing. This approach achieves high-efficiency multiplex editing without DSBs and their associated complications, such as chromosomal translocations and hindered cell proliferation 15. Previously, our group established a platform for the multiplex engineering of human lymphocytes using CRISPR-Cas9 base editors. Concurrent nicking of the unedited strand by the core Cas9 nickase then stimulates DNA repair to use the newly deaminated base as a template for DNA polymerization, thereby preserving the edit in both strands of the DNA. Once bound, the Cas9 complexes displaces the nonbound strand, forming a ssDNA R-loop. The R-loop is rendered accessible to the tethered deaminase domain, whereby cytidine deaminase base editors (CBEs, C:G-to-T:A) deaminate C-to-U, which base pairs like T, and adenosine deaminase base editors (ABEs, A:T-to-G:C) deaminate A-to-I, which base pairs like G. In principle, base editors localize to a target region in the genome guided by a sgRNA. Base editors are a class of gene editing enzymes that consist of a Cas9 nickase fused to a nucleotide deaminase domain 17, 18, 19. However, as more is understood about the complex genetic circuitry involved in cancer immunosurveillance 16, and enthusiasm increases for generating ‘off the shelf’ CAR T cells 1, 2, 3, 4, 5, 7, 15, it is of increased interest to edit genes in a multiplex setting 8.Īn alternative tool to edit genes without causing DSBs are CRISPR-Cas9 base editors. Concerns arising from DSBs are exacerbated in a multiplex setting, where multiple genes are targeted simultaneously 7, 15. 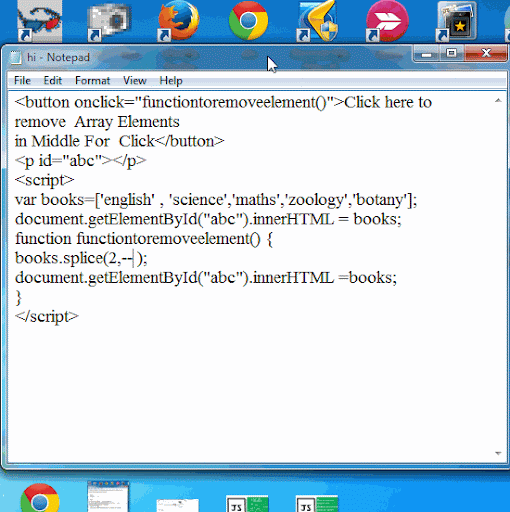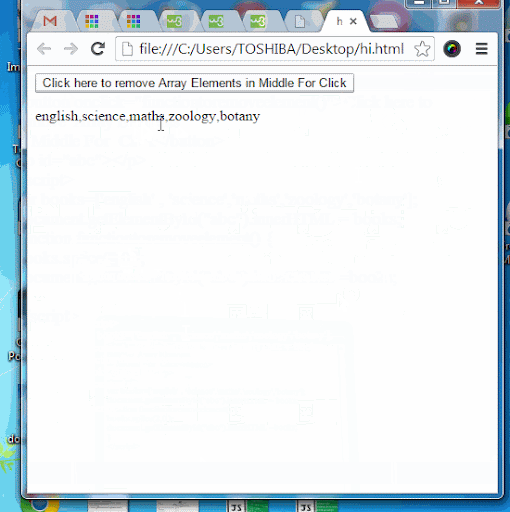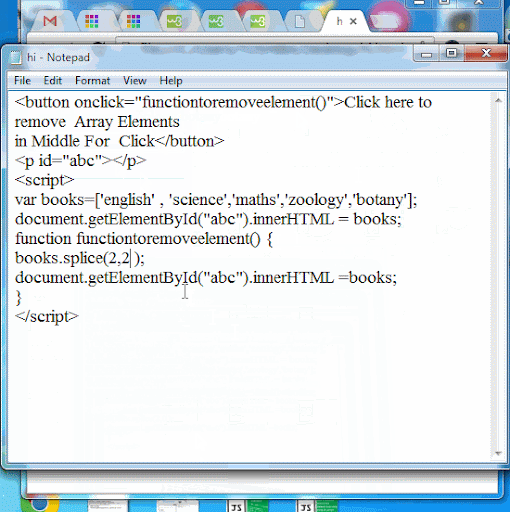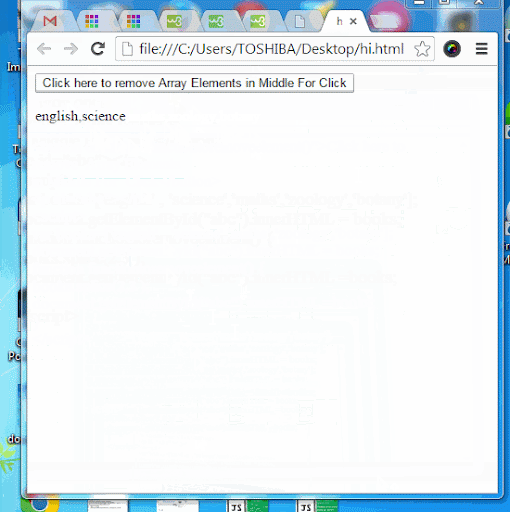
While CRISPR-Cas9 nucleases work exceptionally well for single gene editing, multiple concerns have emerged surrounding DSB induction, including large scale genomic rearrangements 11, 12, loss of heterozygosity 13, and selection of cells with p53 alterations 14, leading to suboptimal efficiency and the potential of oncogenesis in cell based therapies. In the most commonly used form of CRISPR, the Cas9 nuclease from Streptococcus pyogenes (hereafter referred to as Cas9) is paired with a single-guide RNA (sgRNA) to induce a double-stranded break (DSB) at a specific DNA site directed by the programmable complementarity of the 20-nt sgRNA protospacer 5. These applications are of particular interest in the field of cellular immunotherapies, where the multiplexed disruption of genes involved in alloreactivity (e.g., TRAC, TRBC, and B2M) and in immunosuppression (e.g., PDCD1, CTLA4, TGFBR2, and CISH) in tandem with the knockin of chimeric antigen receptors (CARs) specific to tumor antigens yields promise in the development of efficacious and safe therapies to recalcitrant malignancies 7, 8, 9, 10. Key amongst the applications of these systems is their use in gene editing for targeted gene knockout, knockin, and modification 6. Collectively, we demonstrate a robust method for gene disruption, accompanied by a modular design tool that is of use to basic and translational researchers alike.Ĭlustered Regularly Interspaced Short Palindromic Repeats (CRISPR) systems and their CRISPR associated proteins (Cas proteins) have allowed for an unprecedented ability to manipulate the genome 1, 2, 3, 4, 5. Further, the CBE BE4 is more effective for disruption than the ABE ABE7.10, however this disparity is eliminated by employing ABE8e. Among the targeted genes, we find that targeting splice-donors is the most reliable disruption method, followed by targeting splice-acceptors, and introducing pmSTOPs. To address these needs, we develop SpliceR ( ) to design and rank BE-splice sgRNAs for any Ensembl annotated genome, and compared disruption approaches in T cells using a screen against the TCR-CD3 MHC Class I immune synapse.  
However, no in-depth comparison of these methods or a modular tool for designing BE-splice sgRNAs exists. Nature Communications volume 12, Article number: 2437 ( 2021)ĬRISPR-Cas9 cytidine and adenosine base editors (CBEs and ABEs) can disrupt genes without introducing double-stranded breaks by inactivating splice sites (BE-splice) or by introducing premature stop (pmSTOP) codons. CRISPR-Cas9 cytidine and adenosine base editing of splice-sites mediates highly-efficient disruption of proteins in primary and immortalized cells 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
